Hypometric genetics: Improved power in genetic discovery by incorporating quality control flags
Published in The American Journal of Human Genetics, 2024
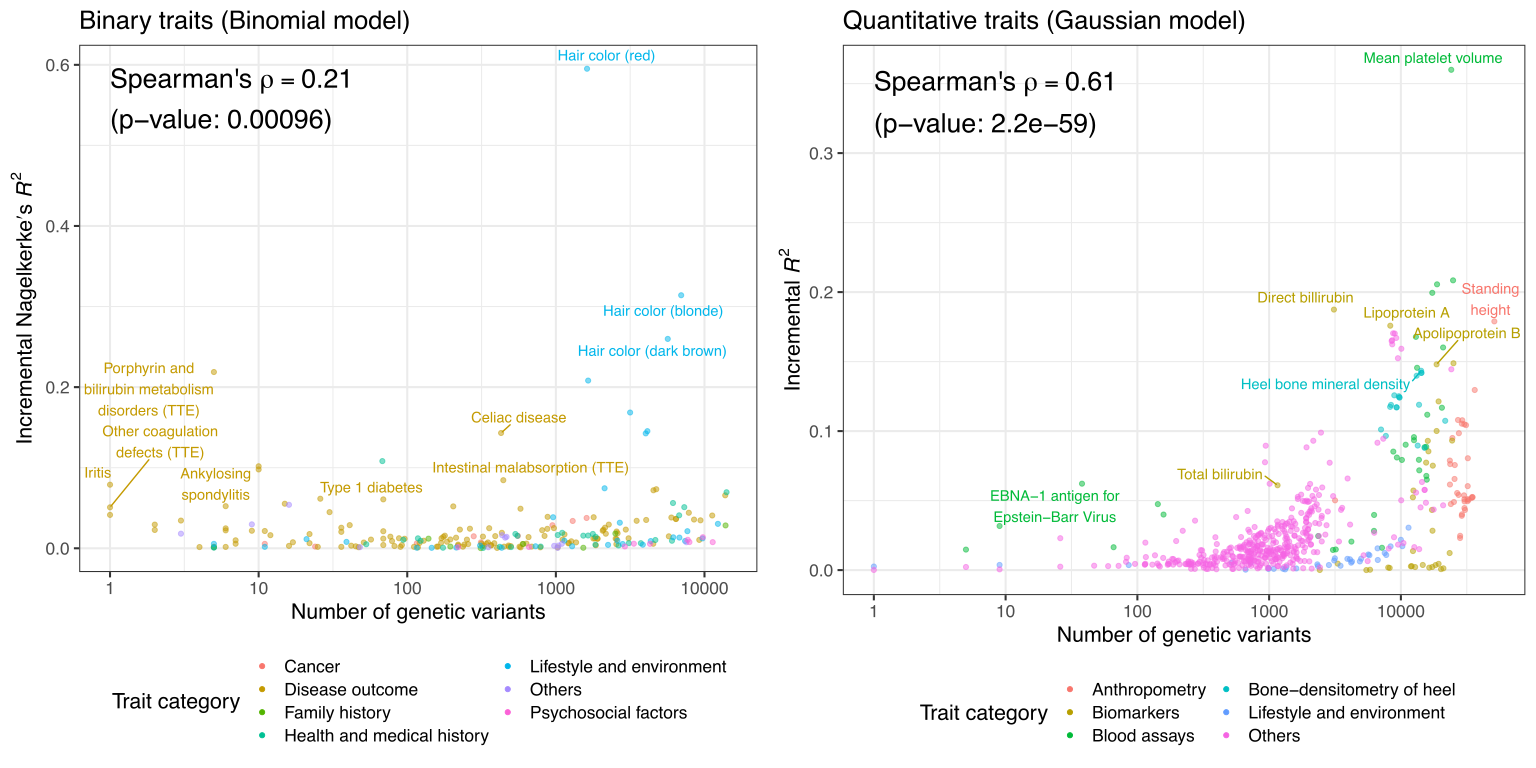
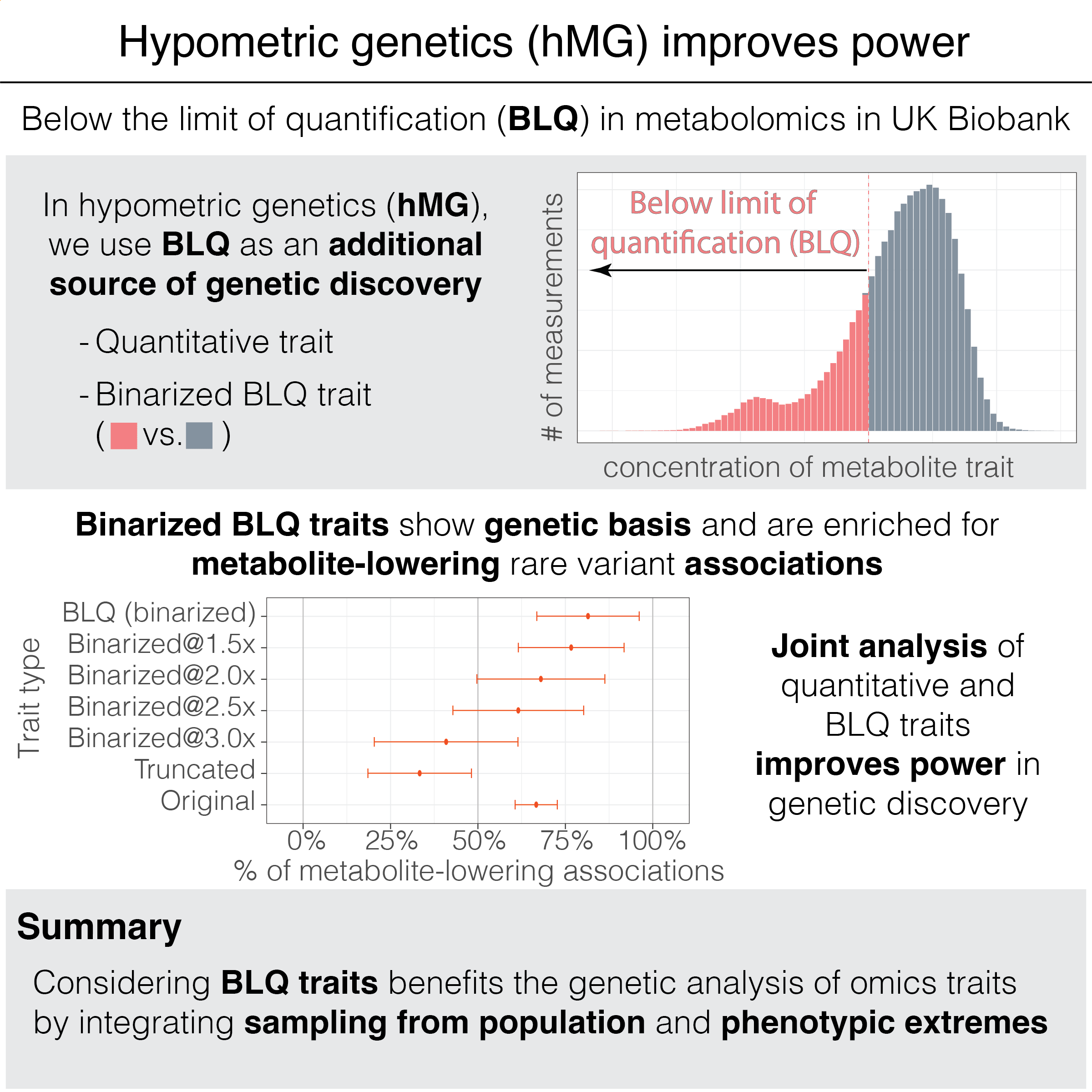
We introduce “hypometric genetics,” an approach to investigate the genetic basis of binarized traits representing the presence of below-the-limit-of-quantification (BLQ) quality control indicators. We show a joint analysis of BLQ and quantitative traits leads to improved power in genetic discovery, highlighting the value of considering population-based samples and phenotypic extremes in genetic studies
Reference: Tanigawa and Kellis. Hypometric genetics: Improved power in genetic discovery by incorporating quality control flags. The American Journal of Human Genetics. 111(11):2478-2493 (2024). https://doi.org/10.1016/j.ajhg.2024.09.008